Two new studies have uncovered important clues about how a prolific pathogen causes disease.
Two new studies have uncovered important clues about how a prolific pathogen causes disease.
Not only does this study suggest potential new targets to counteract disease but it also uncovers new aspects of mammalian cell biology.
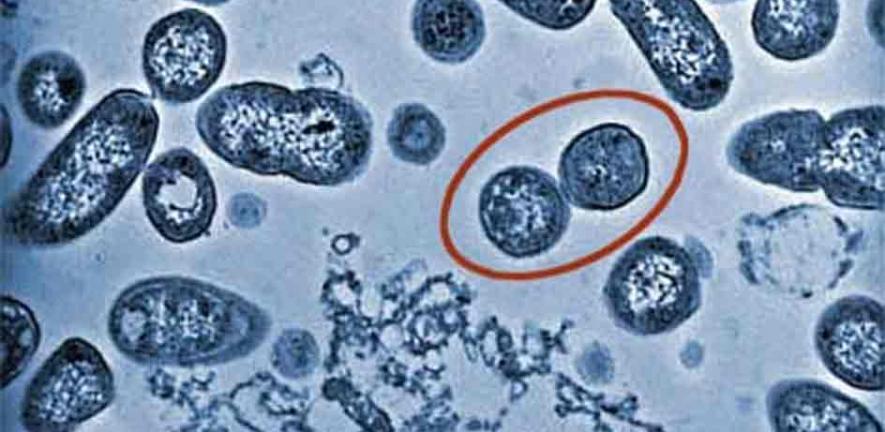
Salmonella are infamous intestinal pathogens that infect humans and animals, causing about 1.3 billion cases of human food-borne diarrhoea and systemic typhoid fever each year throughout the world. Following infection, the bacteria deliver a cocktail of virulence proteins that take control of the cell’s cytoskeleton and networks of signalling pathways, growing inside infected cells in membrane-bound Salmonella-containing vacuoles (SCVs).
Understanding these events at a molecular level has been helped by findings published by Dr Daniel Humphreys and Dr Peter Hume in a team led by Professor Vassilis Koronakis in the Department of Pathology. The research, funded by the Wellcome Trust, has uncovered key features of an unexpected pathway by which Salmonella manipulates the host cell to promote its own growth. The scientists engineered Salmonella to deliver a mutant protein that stalled activity of the protein it binds to in the infected cell, thereby capturing it. Using protein biochemistry and fluorescence microscopy, they showed that the Salmonella protein subverts the assembly of protein complexes required for membrane fusion and formation of intracellular organelles, ensuring a safe, rich, intracellular replicative niche for itself. Not only does this study suggest potential new targets to counteract disease but it also uncovers new aspects of mammalian cell biology.
Meanwhile, scientists at the Department of Veterinary Medicine have pioneered a technique to track the pattern and spread of bacterial infection within the body. The team led by Dr Andrew Grant, Dr Pietro Mastroeni and Professor Duncan Maskell developed molecular tags to mark populations of otherwise identical bacteria; in combination with multicolour fluorescence microscopy, these tagged bacteria could then be tracked during an infection. By constructing mathematical models, the team have managed to tease out the sequence of events, providing a more sophisticated picture of how an infectious disease is driven than previously possible. Thanks to recently awarded funding from the Medical Research Council (MRC), they are now ready to exploit these systems to understand the impact of key host and bacterial factors on the pathogenesis of Salmonella at the level of individual bacterial subpopulations. In the long term, this technique will provide a basis for targeting individual bacterial components in vivo with novel drugs and vaccines that are directed specifically to the sites of infection.
For more information, please contact Professor Vassilis Koronakis (vk103@cam.ac.uk) and Dr Andrew Grant (ajg60@cam.ac.uk).
This work is licensed under a Creative Commons Licence. If you use this content on your site please link back to this page.





