A new technique which traps light at the nanoscale to enable real-time monitoring of individual molecules bending and flexing may aid in our understanding of how changes within a cell can lead to diseases such as cancer.
A new technique which traps light at the nanoscale to enable real-time monitoring of individual molecules bending and flexing may aid in our understanding of how changes within a cell can lead to diseases such as cancer.
It’s like having an extremely powerful magnifying glass made out of gold
Jeremy Baumberg
A new method which uses tightly confined light trapped between gold mirrors a billionth of a metre apart to watch molecules ‘dancing’ in real time could help researchers uncover many of the cell processes that are essential to all life, and how small changes to these processes can lead to diseases such as cancer or Alzheimer’s.
Researchers from the University of Cambridge have demonstrated how to use light to view individual molecules bending and flexing as they move through a model cell membrane, in order to better understand the inner workings of cells. Details are published today (12 August) in the journal Scientific Reports.
The membrane is vital to the normal functioning of cells; keeping viruses out but allowing select molecules, such as drugs, to get through. This critical front line of cellular defence is made up of a layer of fatty lipids, just a few nanometres (one billionth of a metre) thick.
When the cell membrane is damaged however, unwanted invaders can march into the cell. Many degenerative diseases, such as Alzheimer’s, Parkinson’s, cystic fibrosis and muscular dystrophy are believed to originate from damage to the cell membrane.
The ability to watch how individual lipid molecules interact with their environment can help researchers understand not only how these and other diseases behave at their earliest stages, but also many of the fundamental biological processes which are key to all life.
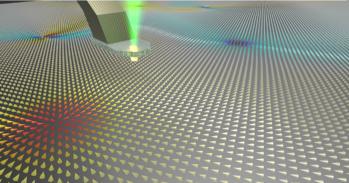
In order to view the behaviour of the cell membrane at the level of individual molecules, the Cambridge team, working with researchers from the University of Leeds, squeezed them into a tiny gap between the mirrored gold facets of a nanoparticle sitting just above a flat gold surface.
Through highly precise control of the geometry of the nanostructures, and using Raman spectroscopy, an ultra-sensitive molecular identification technique, the light can be trapped between the mirrors, allowing the researchers to ‘fingerprint’ individual molecules. “It’s like having an extremely powerful magnifying glass made out of gold,” said Professor Jeremy Baumberg of the NanoPhotonics Centre at Cambridge’s Cavendish Laboratory, who led the research.
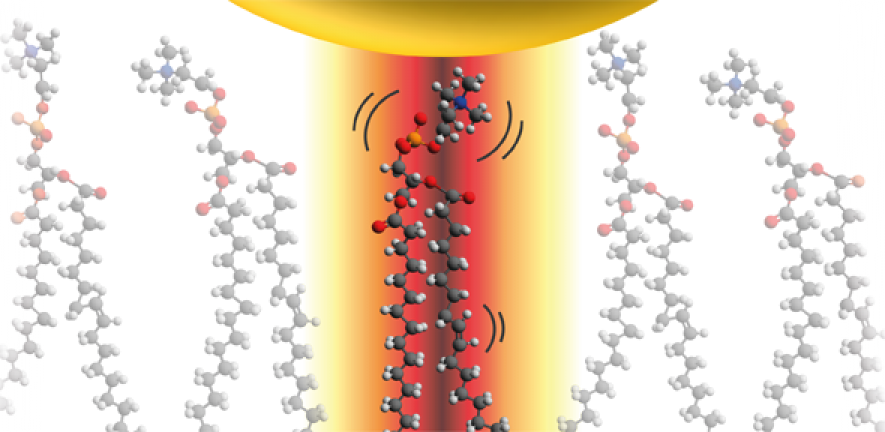
Analysing the colours of the light which is scattered by the mirrors allowed the different vibrations of each molecule to be seen within this intense optical field. “Probing such delicate biological samples with light allows us to watch these dancing molecules for hours without changing or destroying them,” said co-author Felix Benz. The molecules stand shoulder to shoulder like trees in a forest, while a few jitter around sideways.
By continuously observing the scattered light, individual molecules are seen moving in and out of the tiny gaps between the mirrors. Carefully analysis of the signatures from different parts of each molecule allowed any changes in the molecule shape to be observed, which helps to understand how their reaction sites can be uncovered when they are at work. Most excitingly the team says these flexing and bending motions are not expected to occur at the slow time scales of the experiment, allowing the researchers to make videos of their progress.
“It is completely astonishing to watch the molecules change shape in real time,” said Richard Taylor, lead author of the paper.
The new insights from this work suggest ways to unveil processes which are essential to all life and understand how small changes to these processes can cause disease.
The research was funded by the UK Engineering and Physical Sciences Research Council and the European Research Council.
The text in this work is licensed under a Creative Commons Licence. If you use this content on your site please link back to this page. For image rights, please see the credits associated with each individual image.





