JOURNEYS OF DISCOVERY
David Klenerman and Shankar Balasubramanian talk about their discovery of a revolutionary DNA sequencing technology – and the global impact that continues to surprise them.

Impact at a glance
Solexa-Illumina sequencing technology is used to sequence genomes on a population scale, with impact across health and medicine, crop sciences, environmental sciences, bioenergy and more.
During the pandemic, Illumina sequencing underpinned surveillance of the coronavirus worldwide, and also provided sequencing of 35,000 human genomes as part of a UK-wide study of COVID-19.
The technology underpins major genome projects such as the 100,000 Genomes Project, International Cancer Genome Project and Genome Asia 100k.
A whole new sector of the pharmaceutical industry has formed around one of the latest uses of Solexa sequencing technology: detecting ‘cancer signatures’ in pieces of DNA floating around freely in the blood.
The Guinness Book of Records lists the time of 20 hr 10 min as the fastest genetic diagnosis when a team of US doctors read three billion bases of a child’s genome, plus the genomes of both parents, and identified the error causing their rare genetic disorder using an Illumina DNA sequencer.
The benefits to society of rapid genome sequencing are huge. Coronavirus is tracked worldwide, diseases are diagnosed, crops are improved, and new therapies and vaccines are developed. We talk to the inventors about the journey of discovery that began in Cambridge on 26 August 1997.
Shankar Balasubramanian’s diary records the date as the day of “The Solexa Idea!” Sitting in the beer garden of Cambridge’s Panton Arms*, he and David Klenerman sketched out their plans to watch DNA polymerase as it assembled the building blocks of life. Their ideas were progressing fast – and with them, something even more exciting.
Later, he would describe the evening as when “the pieces of the jigsaw came together”. They realised that if they could watch the enzyme copying a genome then they were inadvertently also reading the genome. They had discovered a radically new way to sequence DNA.
But it was when they calculated their possible ‘reading speed’ that it occurred to them they could be looking at something revolutionary: genome sequencing that would be fast, accurate, low-cost and scalable. Within a year, they had co-founded the company Solexa to make the technology more broadly available to the world.
Their hopes for the technology have been exceeded over and over again. Illumina acquired Solexa in 2007 and today Solexa-Illumina ‘next-generation sequencing’ is thought to be responsible for as much as 90% of the total DNA and RNA sequenced in the world, bringing huge benefits to society.
On 18 May 2021, David and Shankar were awarded the Millennium Technology Prize, one of the world’s most prestigious science and technology prizes, by Technology Academy Finland.
On 9 September 2021, the pair were awarded the 2022 Breakthrough Prize in Life Sciences – the world’s largest science prize – for the development of next-generation DNA sequencing.
We talk to the inventors about their journey of discovery.
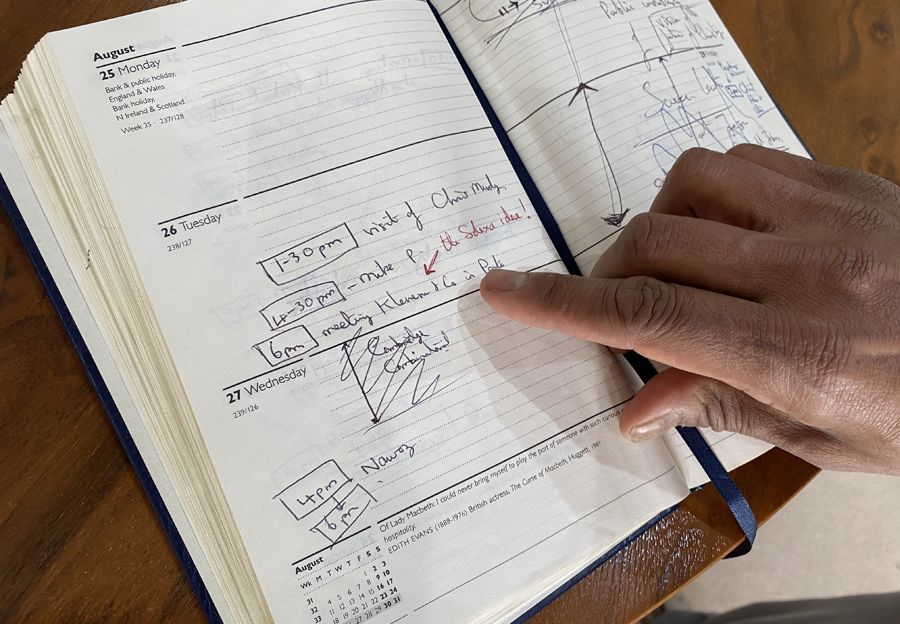
Shankar looks back at his diary for 1997, where he later wrote 'the Solexa idea!' next to 'meeting Klenerman & Co' in the Panton Arms.
Shankar looks back at his diary for 1997, where he later wrote 'the Solexa idea!' next to 'meeting Klenerman & Co' in the Panton Arms.
“It really hit home when I heard someone had used our technology to sequence and treat a baby born with a rare disease. I realised, wow, it’s not just a technology for scientists, it’s actually a technology that can make a difference.”
“The impact and breadth of utility of this technology has gone way beyond my imagination, and it’s still in its infancy.”
Has there been a defining moment for you in seeing the benefits of next-generation DNA sequencing?
DK: “No one knew how much being able to read the building blocks of life would contribute to science and medicine. It really hit home when I heard someone had used our technology to sequence and treat a baby born with a rare disease. I realised, wow, it’s not just a technology for scientists, it’s actually a technology that can make a difference .”
SB: “Probably the biggest surprise to me has been the broad impact of the technology in many unanticipated areas, and this is still on a rising curve. One that I'll single out is the impact of DNA sequencing during the COVID pandemic to track the virus and variants – notably by COG-UK which involves Cambridge University and the Sanger Institute. It's also being used to sequence the genomes of people with COVID-19 to understand why some suffer serious disease while the majority has a benign infection.”
You had an idea to build a machine that could sequence a billion bases of DNA 10,000 times faster than other technology. Why was speed so crucial?
SB: “In the 1990s, a global effort was going into sequencing one genome through the Human Genome Project, but given the goal was to understand the genetic bases of human variation and diseases, then more genomes would need to be sequenced, and ultimately at population scale. We spoke to colleagues at the Sanger involved in the Human Genome Project and asked the question ‘is this the right time for a sequencing technology like ours?’ They were tremendously enthusiastic for us to make it happen.”
DK: “I wrote down a very simple calculation on a flight to Germany and left it in Shankar's pigeon hole. Our proposal was to anchor one strand of DNA to a surface and use DNA polymerase to build a new copy using coloured building blocks that we could measure. The key was to make this massively parallel. It was quite a trivial calculation really, but seeing the answer was the moment I thought ‘this is something’.”
How close were your back-of-the-envelope calculations to what’s happened since?
SB: “Someone once asked if we were ‘sequencing jocks’ back then – we weren’t, we worked in fundamental chemistry. But one August evening in the Panton Arms, we suddenly saw the pieces of the jigsaw come together that led us to see a way of sequencing DNA that could be scalable, massively scalable.”
“We thought it would be ten thousand times faster. It’s actually ten million times faster. It has gone way beyond our expectations.”
DK: “We did some proof-of-concept experiments to check it was possible. We also knew that to achieve what we wanted we had to raise some investment. We went through about nine months of due diligence – this was potentially so revolutionary, potentially so risky. Abingworth decided to fund us at a very modest level with two post-docs in the Chemistry Department. We built up from there, founding Solexa.”
SB: “A big inflexion point was when we built and shipped our first sequencer in 2006, the 1G Genome Analyser. It proved our initial calculations which predicted that one day we would sequence a billion bases.
Illumina became interested in collaborating with Solexa and they acquired the company and the technology in 2007. What they did was take that technology and improve it further by another ten-thousand fold, which is remarkable.
Today, the highest capacity sequencing machine on the planet, Illumina’s NovaSeq 6000, can sequence 48 human genomes (each at 30-fold depth) in just two days; that’s one high-quality genome an hour. I understand that the next-generation sequencing process we started is now responsible for as much as 90% of the total DNA and RNA that gets sequenced in the world.”
Being able to sequence a genome for $100 is seen as the next big milestone in DNA sequencing. Why is this?
SB: “If the cost comes down to $100 a genome, it becomes economically viable to sequence a population and it also becomes practical to sequence people regularly as part of blood-based preventative health checks.”
DK: “To put this in context, the international Human Genome Project started in 1990 and completed in 2003 and cost more than a billion dollars. Of course, that was the very first genome to be sequenced, which makes it much harder to piece together. Today, the human genome can be sequenced in less than one day at a cost of less than $1,000. This has effectively ‘democratised’ sequencing to the point that even quite small labs can have a bench-top DNA sequencer.”
SB: “This democratisation is broadening the impact of sequencing across all fields. I’d never have dreamt of some of the novel applications we’re seeing as a result. That’s what happens when you put technology into the hands of smart, creative researchers worldwide. But the impact and breadth of utility of this technology has gone way beyond my imagination, and it’s still in its infancy.”
What would you say to an early-career researcher with a brilliant idea?
SB: “I say to my students: ‘Don’t think small, think big, and be patient.’ Did everything work first for us time? Absolutely not. It’s essential to fail, that’s how you learn, but try to fail quickly. And take the time to dream how the world might be changed as a result of your idea.”
DK: “You only get opportunities for discovery and innovation at best once or twice in your career. Recognise the moment and do everything you can to get the idea to work. What surprised me was we literally went from a piece of paper to something that was used all over the world in just under 10 years. I wouldn’t have expected things to move that fast and have so much impact in such a short length of time.”
Any last thoughts on the discovery?
SB: “We’re now in a position to start integrating this technology and the information that it generates into national healthcare systems. Linking to the work that David and I did, it all began with a taxpayer-funded research grant here in the UK and without that funding none of this would have happened.”

Credit: Millennium Technology Prize
Credit: Millennium Technology Prize
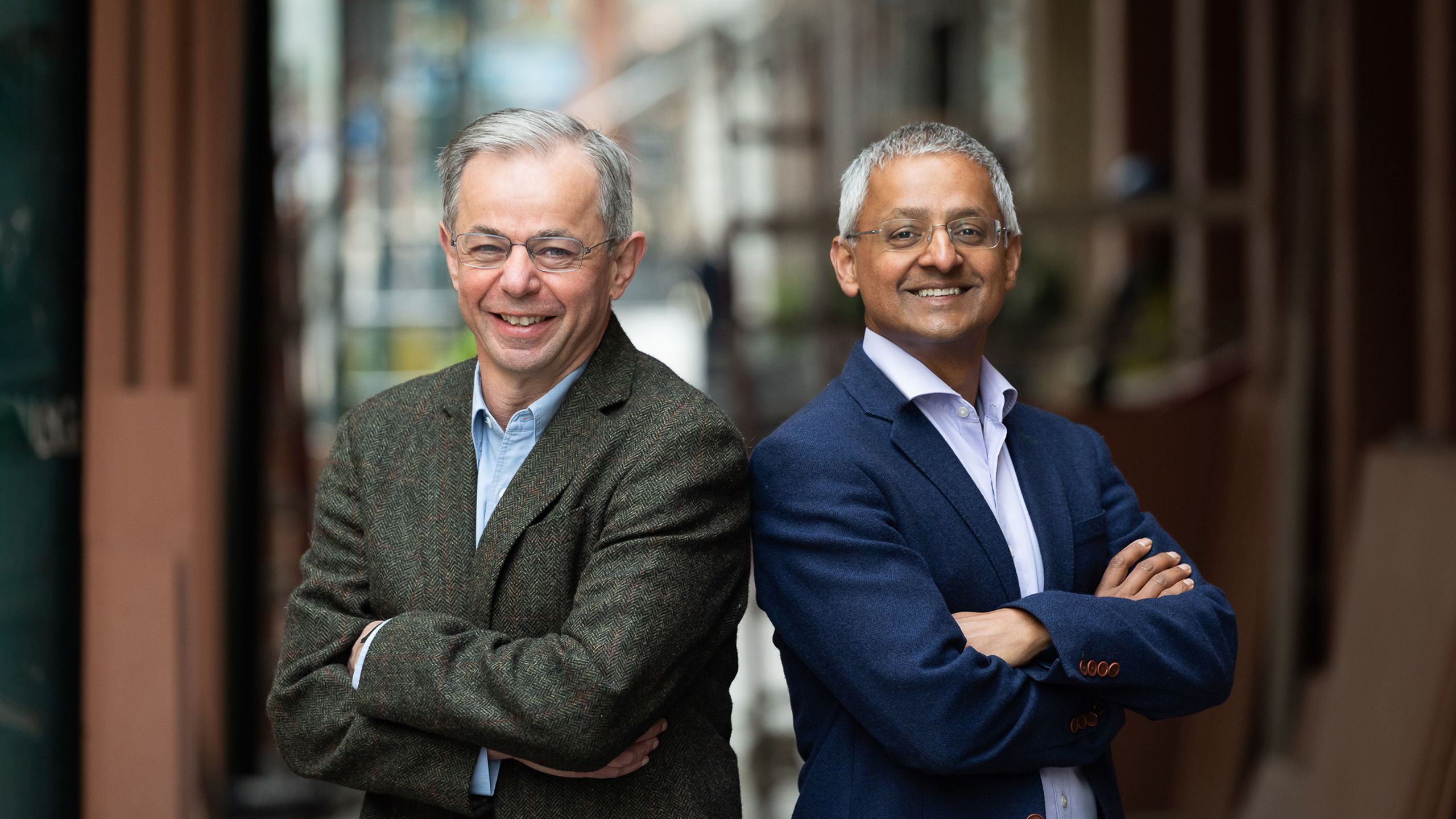
Professor Sir David Klenerman (left) and Sir Shankar Balasubramanian are in the Yusuf Hamied Department of Chemistry; Professor Balasubramanian also has a laboratory at the Cancer Research UK, Cambridge Institute. In 1998, they approached the venture capital firm Abingworth Management and obtained initial seed funding to form Solexa, which was acquired by Illumina in 2007. Since then, llumina has dominated the next generation sequencing market, developing technologies and platforms that radically reduce the run time and cost of Solexa-Illumina Sequencing. Balasubramanian is a Fellow of Trinity College. Klenerman is a Fellow of Christ's College.

How it works
DNA is fragmented into many small pieces that are immobilised on the surface of a chip and locally amplified. Each fragment is then decoded on the chip, base-by-base, using fluorescently coloured nucleotides added by an enzyme.
By detecting the colour-coded nucleotides incorporated at each position on the chip with a fluorescence detector – and repeating this cycle hundreds of times – it is possible to determine the DNA sequence of each fragment.
The collected data is then analysed using sophisticated computer software to assemble the full DNA sequence from the sequence of all these fragments. The method’s ability to sequence billions of fragments in a parallel fashion makes the technique super-fast, accurate and cost-efficient.
*Meanwhile, at the Panton Arms...
“I once told the landlord that I’ll make him very famous one day,” says Shankar.
“I remember going home that August evening feeling pretty excited, as I often did after a beer summit at the Panton Arms. The acid test, of course, is how you feel about it the next day.”
Eleven years later, in November 2008, the Panton Arms patrons were among 100 authors on a paper in Nature describing the sequencing of the first African genome using their sequencing technology.
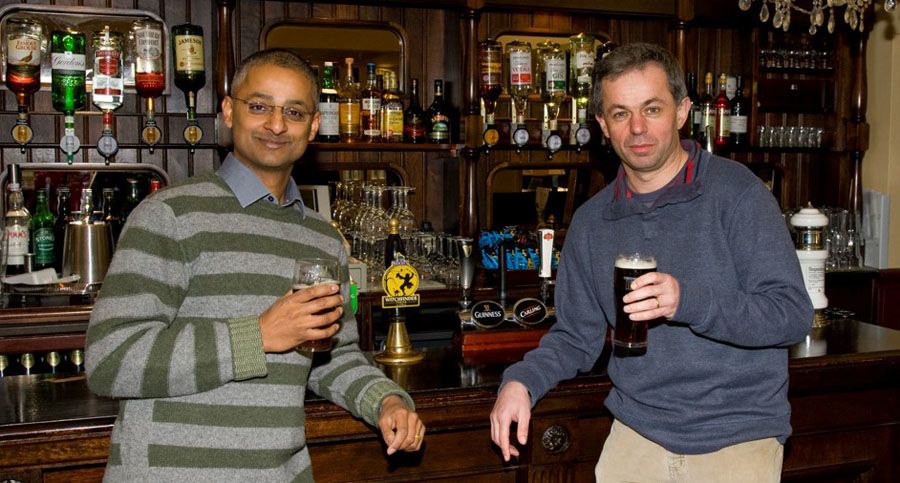

Shankar and David in the Panton Arms (credit: John Holman)
Shankar and David in the Panton Arms (credit: John Holman)

... visit our interactive map to find out how.