“We’re in it for the long haul”
The Cambridge institute taking on COVID-19

In late 2019, a new institute opened on the Cambridge Biomedical Campus. Its timing could not have been better - as the COVID-19 pandemic sent Britain into lockdown several months later, the institute found itself at the heart of the University’s response to this unprecedented challenge.
When I first meet Professor Ken Smith in his new office, out of instinct we go to shake hands. There’s an awkward moment as we remember the public health advice and stop ourselves, opting for ‘elbow bumps’ instead.
It’s 10th March 2020 and I'm at the Cambridge Institute of Therapeutic Immunology and Infectious Disease (CITIID), which opened in autumn 2019 in the Jeffrey Cheah Biomedical Centre with Smith at its Director. We are meeting to discuss his upcoming Nature paper, the first major publication from the institute, which looks at a rare immune disorder known as primary immunodeficiency.
Less than a fortnight later, Britain entered lockdown as SARS-CoV-2 swept the country. Although I didn’t realise it at the time, CITIID would become the last University building I would enter for over three months.
By the time I return, the Jeffrey Cheah building – and, indeed, the world – is a very different place. The bustle on the ground floor has gone – there are far fewer people around and the café is closed. Hand sanitiser dispensers greet you at the entrance and signs on the walls remind you to keep your distance. Stickers in the lift tell you where to stand and that only two people should ride at any one time.
But beyond the visible transformation, something fundamental has changed.
CITIID was one of the few University buildings to remain open throughout lockdown. That’s because, as it became clear the world was facing an unprecedented threat, Smith realised that his team had vital expertise that could help fight the pandemic.

Ken Smith (Credit: Nick Saffell)
Ken Smith (Credit: Nick Saffell)
“We realised we had an institute that was quite consciously set up to deal with this sort of thing,” says Smith. He had intentionally recruited people with interests including immunology, inflammation, infectious disease, virology, cell biology, global health, pathogen surveillance... “We've got this broad range of expertise that allows us to look at the problem from all angles.”
Smith estimates that the institute was running at about 50% capacity, with around 150 scientists, including some from labs elsewhere that had gone into lockdown, but whose work was relevant to the pandemic. Some 60% of the institute’s group leaders also work at one of the hospitals on the Cambridge Biomedical Campus, including Addenbrooke’s Hospital, part of Cambridge University Hospitals NHS Foundation Trust (CUH).
In fact, while CITIID may have been at the heart of the University's response from a biomedical perspective, it’s been a hugely collaborative effort, with scientists from all across the Campus – from other University departments, the Cancer Research UK Cambridge Institute and the Medical Research Council’s Laboratory for Molecular Biology pulling together to fight COVID-19.
Back to basics
Inside CITIID sits the UK’s largest academic Containment Level 3 Facility – the second highest biosecurity level. The facility was scheduled to open in the summer, but through an almost Herculean effort opened several months early. It is essential for enabling researchers to explore the biology of SARS-CoV-2 by culturing live virus.
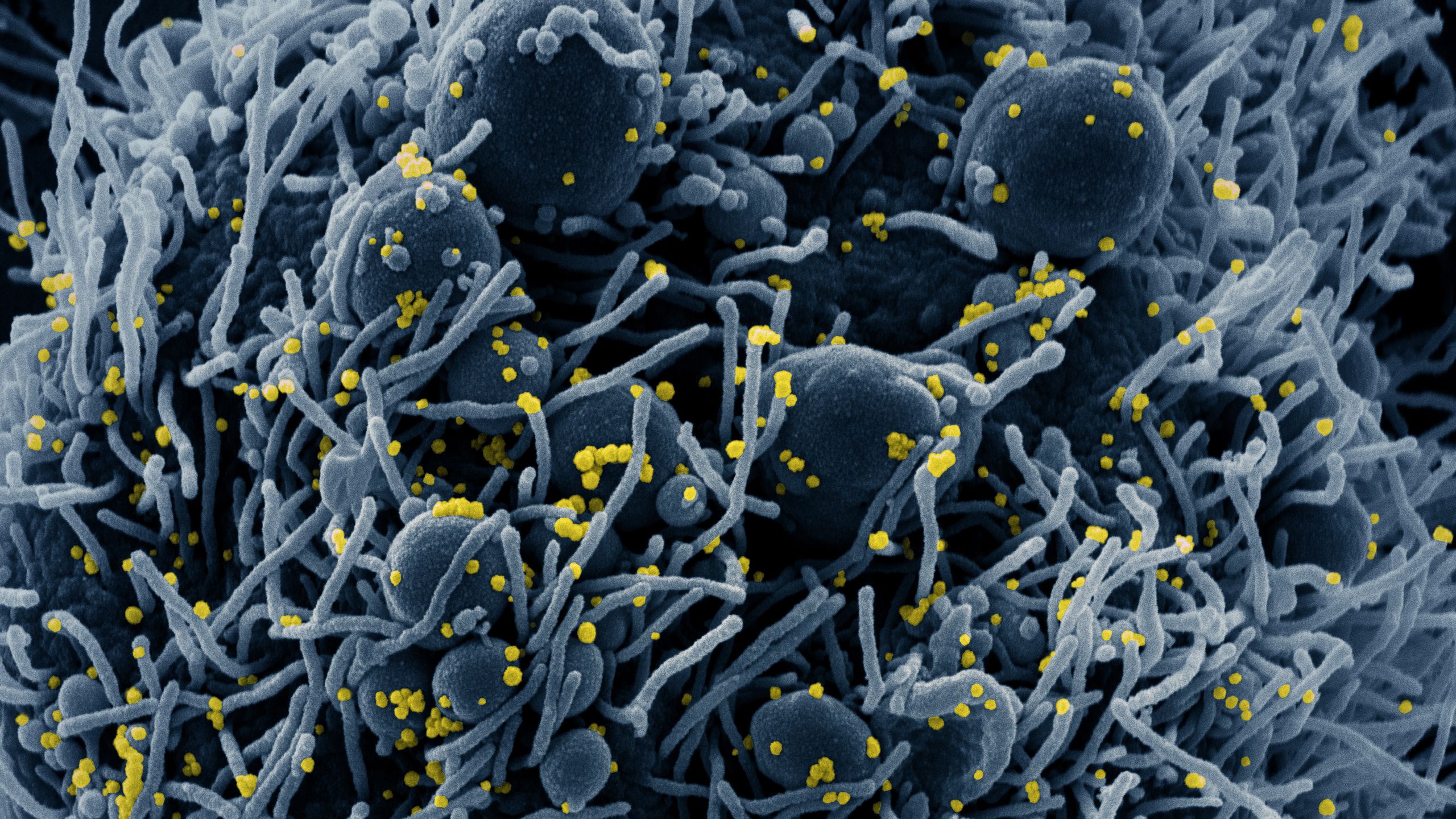
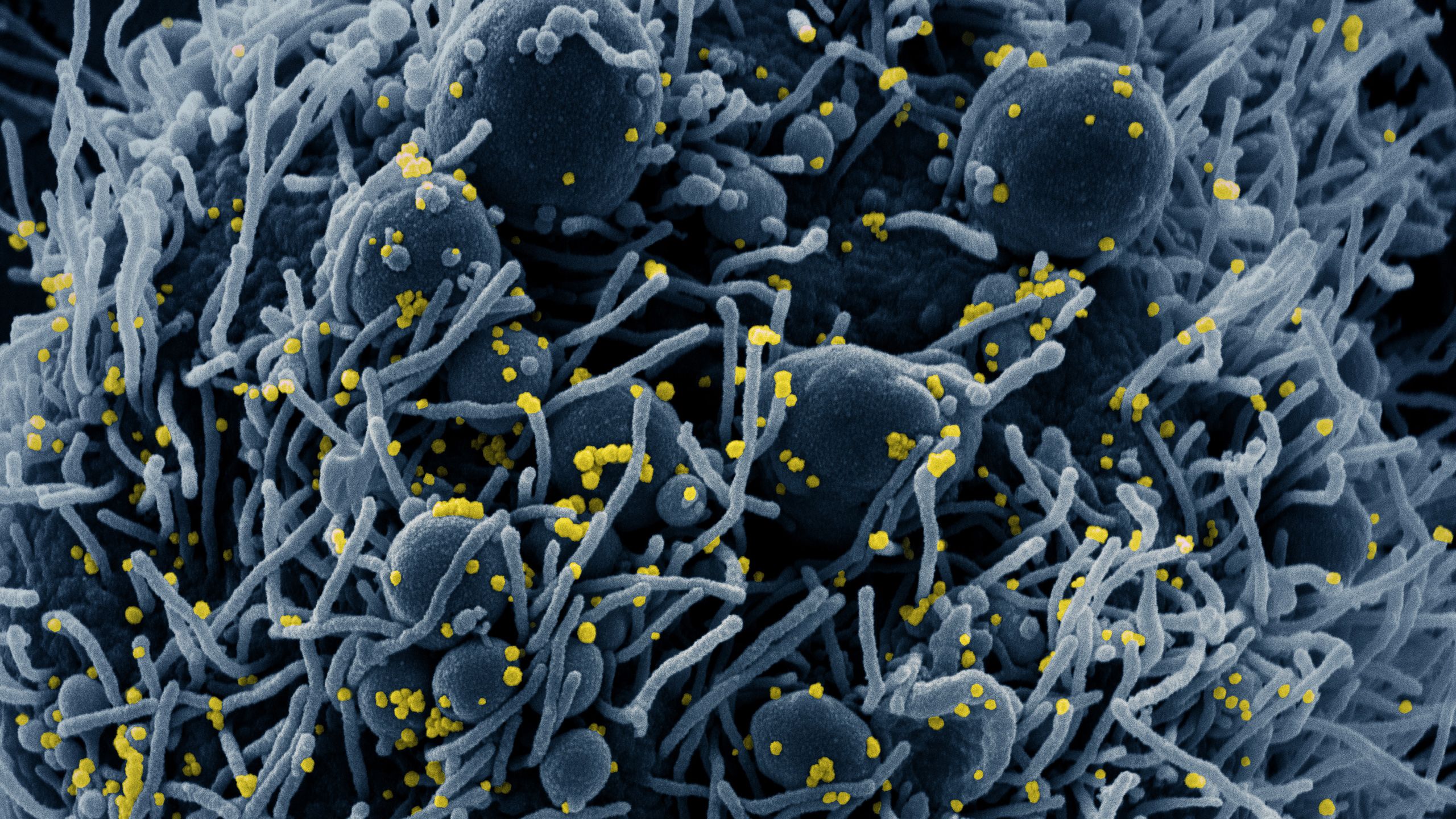
SARS-CoV-2 is a coronavirus, so-named because of its appearance: it is spherical, with ‘spike’ proteins on its surface giving it a crown-like (‘corona’) appearance. This spike protein binds to ACE2, a receptor found on cells in the upper respiratory tract, nasal pharynx and lung – hence why the virus can spread by airborne infection. (ACE2 is also found on cells in the intestine, which may explain why some people experience gastrointestinal symptoms.)
“Despite what you might read, I don't think we know what ACE2 actually does in the airways,” says Professor Paul Lehner, an Infectious Disease Physician who leads CITIID’s Intracellular Immunity Team.
His group is using genetic approaches to understand why ACE2 is important and to look for other factors that might enable the virus to enter cells. They are also using their expertise in proteomics – the study of proteins – to understand how the virus, once inside the cell, manipulates the host cell’s machinery to enable it to replicate, while also evading the immune system.
Lehner is playing the long game: his work may not produce any ‘quick wins’ in the fight against SARS-CoV-2, but that is not the point.
“We’re in it for the long haul,” he says. “What this pandemic has taught us is that we have to be prepared for not just this virus, but the next ones which are coming along. If we can identify new drug targets, which we could use as antivirals, then that might help us with whatever may be coming down the road.”
Smith, too, is interested in the immune system, and in particular the role it plays in the severity of COVID-19 – why is it, for instance, that the disease kills some people while others will not even realise they have been infected?
Smith has teamed up with Professor John Bradley and colleagues at the National Institute for Health Research (NIHR) Cambridge Biomedical Research Centre, to establish the NIHR COVID BioResource. This builds on the existing national NIHR BioResource, initially set up in Cambridge, which collects blood samples from healthy volunteers and patients to examine the links between our genes, environment and health. This meant that when it came to recruiting COVID-19 patients, the infrastructure for collecting the samples was already in place and a legal and regulatory framework existed that could easily be adapted. The samples and associated data will be available to researchers both in Cambridge and across the country.
A team of volunteers from CITIID and other laboratories across Cambridge, together with medical and nursing staff from Cambridge University Hospitals, helped recruit patients, ferry blood samples across the campus and process them, working seven days a week for the first month. “There was a real esprit de corps among the people working on it,” says Smith.
Smith’s work – made possible thanks to this “small army of volunteers” – involves detailed analysis on the samples of exactly which immune cells are present and how this changes over time (a technique known as ‘immunophenotyping’). By marrying this information up against the patient’s medical records, it is possible to see how the symptoms and severity of COVID-19 are related to different changes in the immune system.
“The aim is to identify pathways that are abnormal that we might be able to drug, as some of these might be targets of existing therapeutics,” he explains. “But also, by looking at people with mild disease, can might identify those who have features to suggest they're going to go on to get severe disease. The ability to predict disease outcome is something we need to have, as early treatment is likely to be more effective.”
BBC Look East's Richard Westcott interviews Ken Smith and colleagues about the NIHR COVID Bioresource study.
In the space of just a few months, Smith’s team has managed to create one of the largest, and most intensively immunophenotyped cohorts in the country.
“We've got a terrific cohort,” he says. They have recruited over 200 patients, not only taking blood samples from them throughout their stay in hospital, but also following them up one, three, six and twelve months later. In addition to patients admitted to hospital, the team has recruited health care workers who screened positive for COVID-19, allowing study of those with asymptomatic and mild disease alongside more severe patients.
This is important because one of the more surprising aspects of COVID-19 has been how long recovery can take. Even some patients with only mild disease have reported taking several months before they have fully recovered.
“I think there's a lot more long term damage than we'd envisaged and we just don't know the rate at which the different aspects of the disease improve and correct. We need to follow people up to understand that, and work out ways in which we might help that recovery process. So for example, there may well be long term lung damage that will then put people at risk of subsequent secondary infection, and we need to document that and work out how to manage it."
To achieve this the University and NHS have joined forces to set up multidisciplinary COVID follow-up clinics integrating research and patient care to understand and help manage the long-term effects of the disease.
Another important reason for following patients up over a long people is to measure how long their immune response lasts. Typically, when we catch a new infection, our immune system produces antibodies that will recognise and destroy – or ‘neutralise’ – the infection should we encounter it again, stopping us getting sick a second time. No one knows if this will be the case with SARS-CoV-2 infection.
“Most immune cell types are dramatically reduced in patients with moderate or severe COVID-19, and worryingly some of these do not seem to recover even after some weeks,” says Smith. “The possible long-term consequences of that failure of immune recovery worry us, and is another reason behind our ongoing follow up studies.”
Testing times
As the pandemic started to take hold, Smith recognised that with the NHS facing unprecedented stresses, his team “couldn't ask for the system to do us favours to help research when it was struggling with the capacity to treat sick people” – in fact, he realised his team might actually help relieve some of the pressure on the NHS.
Early on, the team at CITIID, together with the Infectious Diseases team at Addenbrooke’s, established a screening programme for all staff at the hospital, led by Professor Steve Baker and Dr Mike Weekes. This meant frequently testing all healthcare workers – whether symptomatic or not – for SARS-CoV-2 infection. It helped the hospital tighten up its infection control measures – at one point even managing to reduce the number of hospital-acquired infections almost to zero.
The hospital was able to acquire a number of SAMBA II machines developed by Diagnostics for the Real World, a Cambridge spinout company set up by Dr Helen Lee to design monitoring tools for HIV in Africa. These machines enabled rapid point-of-care testing – in other words, they could be deployed in emergency departments, where patients were admitted to the hospital, and provide a fast diagnosis, reducing the time spent in holding areas, releasing valuable single occupancy isolation rooms, and preventing mixing of COVID-19 positive and negative patients.
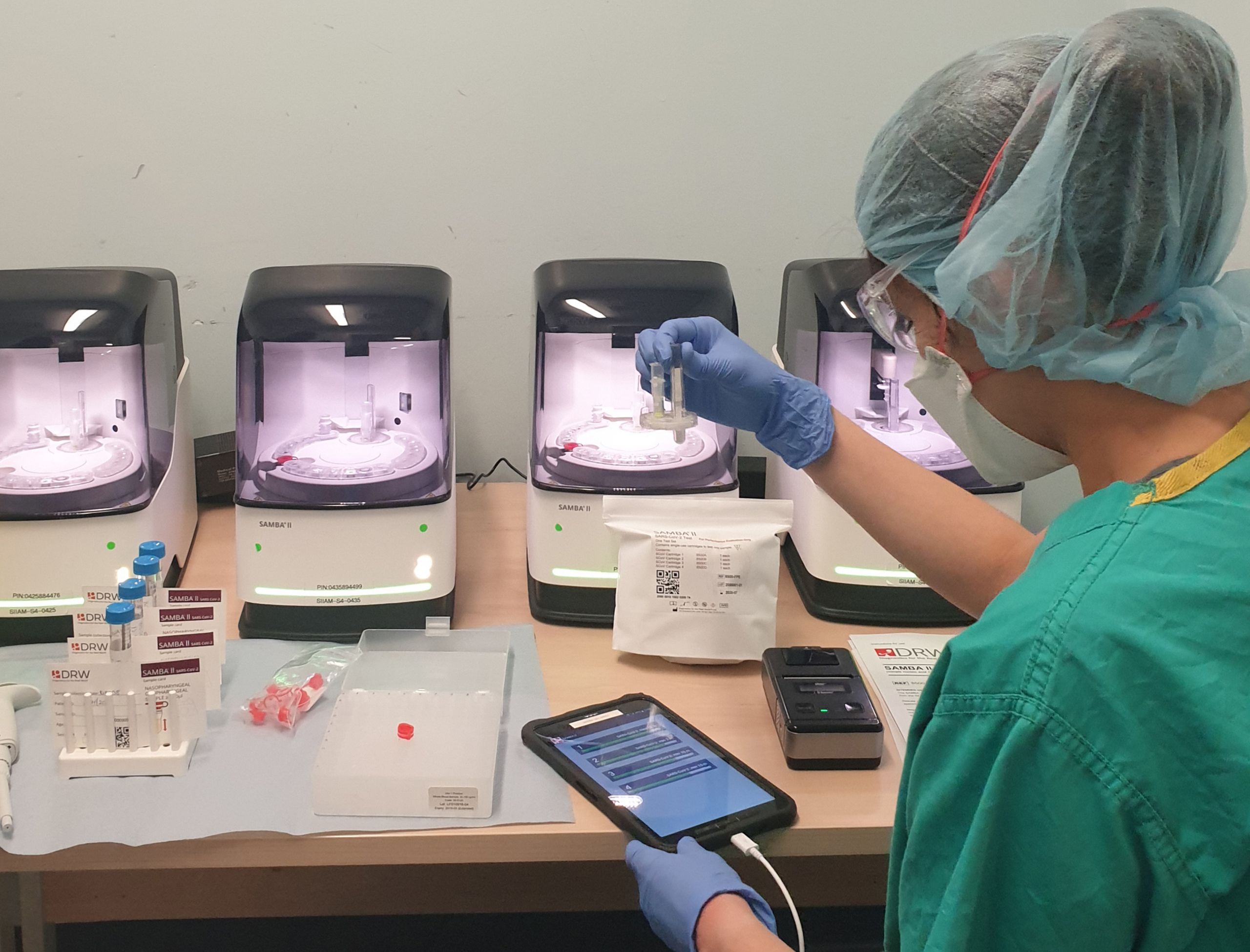
SAMBA II machines at Cambridge University Hospital NHS Foundation Trust
SAMBA II machines at Cambridge University Hospital NHS Foundation Trust
Professor Ravi Gupta, a virologist who works mainly in the HIV field, led an evaluation of the machines and showed that they provided a diagnosis within an average of 2.6 hours compared with 26.4 hours for standard laboratory tests. The average length of time patients had to spend on a COVID-19 ‘holding’ ward before they could be discharged or progress with treatment almost halved: from 58.5 hours to just 30 hours. Gupta’s team found that the testing devices prevented 11 ward closures in the ten days after implementation.
Buoyed by their success, Gupta - who was recently named among TIME magazine's 100 most influential people of 2020 - led a second study which showed that combining the SAMBA tests with antibody tests allowed them to identify 100% of COVID-19 patients, at a time when there was no gold standard test for diagnosing the disease. CUH has since implemented this combination of tests in the emergency department. His team also showed that the rapid antibody tests can detect the mutated SARS-CoV-2 that has taken over from the original strain.

Ravi Gupta (Credit: Nick Saffell)
Ravi Gupta (Credit: Nick Saffell)
Diagnostics for the Real World are currently developing a new version of their SAMBA test capable of detecting both influenza and COVID-19. “That's important because the two diseases are going to look very similar come ‘flu season,” explains Gupta, who hopes to trial these tests, too.
Genomic expertise
For patients unfortunate enough to require hospitalisation, COVID-19 may be just the start of their problems. In very severe disease, patients may require treatment in intensive care units (ICUs) – and at the height of the pandemic, as many as one in three patients admitted to ICU would die.
One of the key reasons, says Professor Gordon Dougan, is secondary infections – while COVID-19 may have made them sick in the first place, they may also be carrying other pathogens such as MRSA and E. coli, which compound their illness.
“When [patients] are put on ventilators, the thing that normally brings them down is not just the COVID, it's the fact that they've got a nasty bacteria lurking there,” he says.
Even before COVID-19, Dougan was already involved in genomic surveillance of antimicrobial resistant pathogens – so-called ‘superbugs’ that can no longer be treated by frontline medicines. His team, along with the ICU staff and Public Health England in Cambridge, have has been carrying our routine testing on patients brought into ICU at CUH.
“What we're able to do it is tell the doctor what that secondary infection is and whether or not they can treat it with a particular antibiotic. We could do that in just a few hours.”

Gordon Dougan (Credit: Nick Saffell)
Gordon Dougan (Credit: Nick Saffell)
Speed of diagnosis is crucial. In most hospitals, it can take several days to get a result, during which time, out of necessity, the patient would have been given a broad-spectrum antibiotic rather than a more targeted antibiotic (or even none, if appropriate).
“This is not good because you breed antibiotic resistance and you can actually make the case worse,” Dougan explains. “Because we can tell [the medics] within a few hours, they'll wait, and then they can give a much more targeted treatment to the patient.”
This approach is proving very successful, with evidence accumulating for better antibiotic management as clinicians can be given much more informed advice.
Because of its expertise in genomic science, Cambridge – the University and nearby Wellcome Sanger Institute – has become the centre of a national initiative, the £20million COVID-19 Genomics UK (COG-UK) Consortium, led by Professor Sharon Peacock, Chair of Public Health and Microbiology at Cambridge and Director of the National Infection Service at Public Health England.
Dougan is part of COG-UK, as is Professor Goodfellow from the Department of Pathology, who is also leading some of the antibody research with Professor Steve Baker at CITIID.
Together with collaborator Dr Estée Török, Goodfellow has shown how genomic surveillance has helped CUH investigate clusters of COVID-19 infections at the hospital. This has led to changes in the hospital’s infection control measures that have helped reduce the number of hospital-acquired infections, making it a safer place for both patients and staff.
Goodfellow, it turns out, is no stranger to epidemics. He has previously spent a large amount of time in Sierra Leone and the Democratic Republic of the Congo helping train local scientists carry out genome sequencing in the ongoing fight against Ebola. Although this experience helped him set up in Cambridge quickly, it also presented a problem: all of his sequencing equipment was on another continent.
Fortunately, the Cambridge community responded quickly, in part because their own labs had closed.
“We had to borrow equipment from people around us,” he explains, “but we were still able to get going within 24 hours and start sequencing the virus. [Our previous work] meant we knew how to do sequencing and we were able to scale up the process. Doing it in Cambridge is much easier.”
The major challenge has been around sample logistics: tracking a sample from the diagnostic lab, through the sequencing and uploading the data, maintaining the link between the patient’s information, and the sequence itself.
As Goodfellow explains, “If you break that connection, the data is worthless.”
"We shouldn't have to rely on luck"
It is sometimes hard to acknowledge that just 12 month ago, no one had heard of SARS-CoV-2 or COVID-19. Now, phrases such as social distancing and lockdown – even the R number – are part of everyday conversations. The coronavirus has changed the world dramatically: will we ever defeat it?
“I think it’s not going to go away in a hurry, that it's going to be an illness that we have in the community for some time. A bit like the flu, it will come and go and be something we have to deal with in an ongoing fashion,” says Smith. “The question is to determine what both society and the medical profession need to do about reducing its impact.”
Even if we cannot eradicate it through vaccination, he believes we will improve its management with antiviral and anti-inflammatory drugs and therapeutics based on antibodies against SARS-CoV-2.
“I think we'll get better at managing severe disease as we understand the disease better. We'll develop better and more specific antivirals, just as we did for HIV. So, even in the absence of a highly effective vaccine, I think it's likely that our medical care will improve. We'll be able to intervene earlier and more effectively and the mortality rate will drop.”
Lehner describes himself as “very optimistic” in the long term, agreeing that effective antivirals and vaccines will help us control the disease. “We've got to get the adult population through this and that's why vaccination is so important, but it might be that in the future we just see it as an infection that we want our kids to get, like chickenpox, or the other circulating, non-pathogenic coronaviruses.”
Prior to COVID-19, experts had long been warning of the risks of the next pandemic. H5N1 bird flu, SARS, MERS and Ebola all threatened to spread beyond regional boundaries, but ultimately failed to become pandemics.
“In a way, we have been relatively lucky so far,” says Dougan. “Even with SARS-CoV-2, this could have been a much more aggressive and virulent virus, for example impacting children.”
Dougan argues that richer nations have gradually lost their sense of danger concerning epidemics and serious infections. “We must reacquire this instinctive memory. We shouldn’t have to rely on luck.”
That’s why having an institute like CITIID – and the huge collaborative effort towards combating COVID-19 among Cambridge’s biomedical departments – has been so important, says Smith.
“What we’re learning about the relationship between infectious disease and our immune systems will help us in this pandemic – and it will also help us to be ready for what comes next.”
To discover more researchers playing a key role in the fight against COVID-19 - including infectious disease expert Dr Estée Török, intensive care specialist Dr Charlotte Summers and statistician Professor Daniela De Angelis - see our series Tackling COVID-19.
Main image: Jeffrey Cheah Biomedical Centre lit up to mark Clap for the Carers (Lloyd Mann)
Sub-header image: Colorized scanning electron micrograph of an apoptotic cell (blue) infected with SARS-COV-2 virus particles (yellow), isolated from a patient sample. Image captured at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. (NIH Image Gallery)